Abstract
Five carbamate derivatives, obtucarbamates C and D (1, 2), dimethyl ((carbonylbis(azanediyl))bis(2-methyl-5,1-phenylene))dicarbamate (3), obtucarbamates A and B (4, 5), and four aromadendrane-type sesquiterpenoids, (+)-4β-N-methenetauryl-10β-methoxy-1β,5α,6β,7β-aromadendrane (6), (−)-4β-N-methenetauryl-10β-methoxy-1β,5β,6α,7α-aromadendrane (7), (−)-4α,10β-aromadendranediol (8), (+)-4β,10β-aromadendranediol (9) were obtained from the South China Sea gorgonian coral Melitodes squamata Nutting. Compounds 1, 2, 6, and 7 were new, and their structures were established by spectroscopic analyses. Compounds 6 and 7 contained a taurine group that was rarely found in marine natural compounds, and 7 showed moderate antibacterial activity. The possible biosynthesis routes of 1–5 were conjectured.
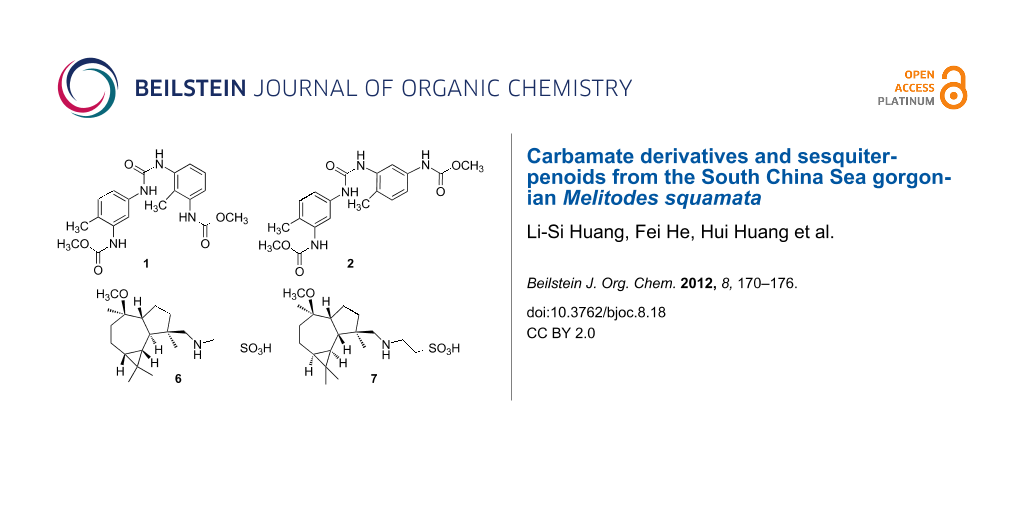
Graphical Abstract
Introduction
Gorgonians are recognized to mainly produce acetogenins, sesquiterpenoids, diterpenoids, prostanoids, and steroids [1,2]. However, nitrogen-containing compounds were relatively few obtained from gorgonians. Gorgonian Melitodes squamata Nutting belonging to Melithaea family is a kind of pharmaceutical coral that has the efficacy of relieving cough, bleeding, and diarrhea, soothing nerves, and calming scare, etc [3]. A previous study on the chemical constituents of Melithaea family gorgonians led to the isolation of four new steroids melithasterols A–D from Melithaea ocracea [4].
Results and Discussion
In order to further obtain new bioactive compounds from gorgonians, we studied the chemical constituents of the South China Sea gorgonian M. squamata, which led to the obtainment of five carbamate derivatives, obtucarbamates C and D (1, 2), ((carbonylbis(azanediyl))bis(2-methyl-5,1-phenylene))dicarbamate (3) [5], obtucarbamate A (4) [6], obtucarbamate B (5) [6], and four aromadendrane-type sesquiterpenoids, (+)-4β-N-methenetauryl-10β-methoxy-1β,5α,6β,7β-aromadendrane (6), (−)-4β-N-methenetauryl-10β-methoxy-1β,5β,6α,7α-aromadendrane (7), (−)-4α,10β-aromadendranediol) (8) [7], (+)-4β,10β-aromadendranediol) (9) [7] (Figure 1) from the EtOH/CH2Cl2 extracts of M. squamata. Compounds 1, 2, 6, and 7 have not been previously reported. The cytotoxicity of 1–9 against human malignant melanoma A735 and cervical carcinoma HeLa cell lines, and the antibacterial activity of 1–5 and 7 towards bacteria Escherichia coli, Bacillus subtilis and Micrococcus luteus were evaluated. The possible biosynthesis routes of 1–5 were conjectured.
Compound 1 showed the same molecular formula of C19H22N4O5 as 3, which was determined by its HRMS–ESI (m/z 409.1568 [M + Na]+) and NMR spectra. The 1H NMR spectrum of 1 (Table 1) exhibited signals for three ABX system aromatic protons at δH 7.50 (s, 1H), 7.19 (dd, J = 2.0, 8.0 Hz, 1H), 7.06 (d, J = 8.0 Hz, 1H), three ABC system aromatic protons at δH 7.57 (d, J = 8.0 Hz, 1H), 7.08 (t, J = 8.0 Hz, 1H), 6.99 (d, J = 8.0 Hz, 1H), two methyl groups at δH 2.12 (s, 3H), 2.07 (s, 3H), and two methoxy groups at δH 3.63 (s, 3H), 3.64 (s, 3H). It also showed four –NH protons at δH 8.90 (s), 8.78 (s) and 8.52 (s, 2H). The 13C NMR and DEPT 135 spectral data (Table 1) showed the presences of 19 carbons, including three carbonyl carbons (δC 154.5, 154.6, 152.7), 12 aromatic carbons, two methyl carbons (δC 16.9, 12.4), and two methoxy carbons at δC 51.5. The NMR data of 1 were very similar to those of compounds 3–5 [5,6]. Compound 3 is a symmetric urea derivative that was formed from the amidation of two molecules of 4 [6]. Comparison of the NMR data of 1 with 3–5 suggested that 1 was an asymmetric urea derivative that was formed from the amidation of 4 and 5 [5,6].
Table 1: 1H (500 MHz) and 13C NMR (125 MHz) data of 1 and 2 (in DMSO-d6, δ in ppm, J in Hz).
| C | 1 | 2 | ||
|---|---|---|---|---|
| δH | δC | δH | δC | |
| 1 | — | 136.5, C | — | 136.3, C |
| 2 | 7.50 (s) | 114.0, CH | 7.54 (s) | 113.8, CH |
| 3 | — | 138.3, C | — | 138.0, C |
| 4 | 7.19 (dd, 2.0, 8.0) | 114.5, CH | 7.19 (br d, 8.0) | 114.0, CH |
| 5 | 7.06 (d, 8.0) | 130.1, CH | 7.06 (d, 8.0) | 130.1, CH |
| 6 | — | 124.3, C | — | 124.0, C |
| 1’ | — | 137.1, C | — | 137.5, C |
| 2’ | — | 124.0, C | 7.93 (s) | 111.4, CH |
| 3’ | — | 137.1, C | — | 137.0, C |
| 4’ | 7.57 (d, 8.0) | 119.2, CH | 7.04 (d, 8.0) | 112.5, CH |
| 5’ | 7.08 (t, 8.0) | 125.2, CH | 7.07 (d, 8.0) | 129.8, CH |
| 6’ | 6.99 (d, 8.0) | 121.8, CH | — | 121.7, C |
| 7–CH3 | 2.12 (s) | 16.9, CH3 | 2.12 (s) | 17.2, CH3 |
| 7’–CH3 | 2.07 (s) | 12.4, CH3 | 2.16 (s) | 16.9, CH3 |
| OCH3 | 3.64 (s) | 51.5, CH3 | 3.64 (s) | 51.5, CH3 |
| 3.63 (s) | 51.5, CH3 | 3.63 (s) | 51.3, CH3 | |
| –NHCONH– |
8.78 (s)
8.90 (s) |
152.7, C |
8.78 (s)
9.46 (s) |
152.5, C |
| –NHCOO– |
8.52 (s)
8.52 (s) |
154.5, C
154.6, C |
8.53 (s)
8.53 (s) |
154.6, C
154.2, C |
The suggestion was supported by the HMBC correlations (Figure 2). In the HMBC spectrum of 1, correlations of Me-7 (s, 2.12) with δC 130.1 (d, C-5), 124.3 (s, C-6), 136.5 (s, C-1) and Me-7’ (s, 2.07) with δC 124.0 (s, C-2’), 137.1 (s, C-1’), 137.1 (s, C-3’) indicate a methyl group attached at both C-6 and C-2’. The HMBC correlations of δH 3.64 (s) with δC 154.5 (s) and δH 3.63 (s) with δC 154.6 (s) revealed the presence of two –NHCOOMe groups. The HMBC correlations of δH 8.78 (NH) with δC 124.3 (C-6), 114.0 (C-2), δH 8.90 (NH) with δC 124.0 (C-2’), 119.2 (C-4’), and δH 8.52 (2NH) with δC 114.0 (C-2), 114.5 (C-4), 121.8 (C-6’), 124.0 (C-2’), suggest that the four –NH groups are attached at C-1, C-3, C-1’, C-3’ of two aromatic rings. Based on the above data analysis, the structure of 1 was elucidated to be as shown above and named as obtucarbamate C.
Compound 2 also exhibited the molecular formula of C19H22N4O5 as deduced from NMR and HRMS–ESI (m/z 409.1576 [M + Na]+). The 1H NMR data of 2 (Table 1) exhibited signals for two ABX aromatic systems, including six aromatic protons at δH 7.54 (s, 1H), 7.19 (br d, J = 8.0 Hz, 1H), 7.06 (d, J = 8.0 Hz, 1H), 7.93 (s, 1H), 7.07 (d, J = 8.0 Hz, 1H), 7.04 (d, J = 8.0 Hz, 1H). The 1H and 13C NMR spectral data of 2 show similarities to those of 3 [5] (Table 1). Comparison of the NMR data of 2 and 3 suggests that 2 is an asymmetric urea derivative that was formed from the amidation of two molecules of 4 [5,6], and the only difference between 2 and 3 is the amidation position of the two molecules of 4. Based on the 1H NMR, 13C NMR HSQC, HMBC and 1H–1H COSY spectral data analysis, the structure of 2 was elucidated to be as shown above and named as obtucarbamate D.
Urea derivatives are closely related in structure to carbamates. Urea is synthesized in the body of many organisms as part of the urea cycle, which is namely a cycle of biochemical reactions occurring in many animals that produces urea ((NH2)2CO) from ammonia (NH3). According to the reactions of the urea cycle [5], we made conjectures about the possible biosynthesis routes of compounds 2–4 as shown (Figure 3). The possible biosynthesis routes to 1, 5 and 2–4 are essentially the same, except starting from 4-methylbenzene-1,3-diamine in place of 2-methybenzene-1,3-diamine. This is the first time that carbamate derivatives from gorgonians have been reported.
Figure 3: Possible biosynthesis routes of compounds 2–4.
Figure 3: Possible biosynthesis routes of compounds 2–4.
Compound 6 was assigned the molecular formula of C19H35NSO4 as deduced from NMR spectra and HRMS–ESI (m/z 372.2170 [M − H]−). The 1H NMR data (Table 2) indicate an aromadendrane skeleton with two cyclopropyl protons resonating at δH 0.44 (t, J = 10.0 Hz, 1H) and 0.67 (ddd, J = 6.5, 10.0, 11.5 Hz, 1H) [7-11]. The 1H NMR spectrum also showed four singlet methyl groups (δH 0.93, 1.02, 1.11, 1.11) and a methoxy group (δH 3.16). The 13C NMR spectrum (Table 2) displayed 19 carbon signals, including five methyl carbons (δC 16.6, 28.7, 17.6, 17.8, 48.3), seven methylene carbons (δC 24.0, 37.0, 19.5, 37.2, 45.8, 45.9, 58.9), four methine carbons (δC 50.1, 46.0, 25.6, 26.6), and three quaternary carbons (δC 19.6, 44.0, 79.1). These NMR data (Table 2) showed similarities to those of (–)-4α,10β-aromadendranediol (8), (+)-4β,10β-aromadendranediol (9) and other analogues [7-11], which suggests that 6 is an aromadendrane-type sesquiterpenoid with a methoxy group and a side chain that contained three methylene carbons.
Table 2: 1H (500 MHz) and 13C NMR (125 MHz) data of 6 and 7 (in CDCl3, δ in ppm, J in Hz).
| No. | 6 | 7 | ||
|---|---|---|---|---|
| δH | δC | δH | δC | |
| 1 | 2.23 (dd, 8.0, 17.0) | 50.1, CH | 2.09 (m) | 52.2, CH |
| 2 | 1.73 (m) | 24.0, CH2 | 1.67 (m) | 25.2, CH2 |
| 3 |
1.37 (t, 11.0),
1.80 (overlap) |
37.0, CH2 | 1.61 (overlap) | 33.4, CH2 |
| 4 | — | 44.0, C | — | 44.3, C |
| 5 | 0.98 (br t, 10.0) | 46.0, CH | 1.72 (overlap) | 41.7, CH |
| 6 | 0.44 (t, 10.0) | 25.6, CH | 0.22 (t, 9.5) | 24.3, CH |
| 7 | 0.67 (ddd, 6.5, 10.0, 11.5) | 26.6, CH | 0.62 (m) | 28.7, CH |
| 8 |
0.84 (ddd, 6.5, 12.5, 13.5),
1.87 (ddd, 6.5, 11.5,13.5) |
19.5, CH2 |
1.14 (m)
1.59 (m) |
18.2, CH2 |
| 9 |
1.59 (t, 12.5)
1.67 (dd, 6.5, 14.5) |
37.2, CH2 |
1.48 (t, 12.0),
1.83 (dd, 6.5, 14.5) |
32.8, CH2 |
| 10 | — | 79.1, C | — | 79.1, C |
| 11 | — | 19.6, C | — | 19.3, C |
| 12 | 0.93 (s) | 28.7, CH3 | 1.01 (s) | 28.3, CH3 |
| 13 | 1.02 (s) | 16.6, CH3 | 0.99 (s) | 16.0, CH3 |
| 14 | 1.11 (s) | 17.6, CH3 | 1.20 (s) | 21.8, CH3 |
| 15 | 1.03 (s) | 17.8, CH3 | 1.10 (s) | 24.4, CH3 |
| 16 |
2.72 (d, 12.0),
2.94 (d, 12.0) |
58.9, CH2 |
2.86 (d, 12.5),
3.04 (d, 12.5) |
58.3, CH2 |
| 17 | 3.54 (t, 5.0) | 45.8, CH2 | 3.27 (br s) | 45.9, CH2 |
| 18 | 3.31 (t, 5.0) | 45.9, CH2 | 3.44 (br s), 3.53 (br s) | 45.7, CH2 |
| OCH3 | 3.16 (s) | 48.3, CH3 | 3.10 (s) | 48.1, CH3 |
This suggestion was proved by the HMBC correlations (Figure 4). In the HMBC spectrum of 6, correlation of MeO– (δH 3.16, s) with δC 79.1 indicates the methoxygenation of C-10. The HMBC correlations of H-16 [δH 2.72 (d, J = 12 Hz), 2.94 (d, J = 12 Hz)] with C-3 (δC 37.0)/C-4 (δC 44.0)/C-5 (δC 46.0)/Me-14 (δC 17.6) indicate that there is a side chain at C-4 instead of a –OH group. The HMBC spectrum also shows correlations of H-16 with C-17 (δC 45.8) and H-17 [δH 3.54 (t, J = 5.0 Hz)] with C-16 (δC 45.8)/C-18 (δC 45.8). In the 1H–1H COSY spectrum (Figure 4), correlation of H-17 with H-18 [δH 3.31 (t, J = 5.0 Hz)] is observed. The above HMBC and 1H–1H COSY spectral data, combined with the molecular formula of C19H35NSO4, suggest the presence of a –CH2NHCH2CH2SO3H group. The suggestion was supported by the 1H and 13C NMR data comparison of the –CH2NHCH2CH2SO3H group in 6 with literature data [12,13]. The IR spectrum of 6 contains two strong bands at 1217 and 1041 cm−1, which supports the presence of a sulfonic acid group. The –NHCH2CH2SO3H group has rarely been found in marine natural compounds, such as N-methyltaurine, taurine, and spongidine D from sponges [12,13].
![[1860-5397-8-18-4]](/bjoc/content/figures/1860-5397-8-18-4.png?scale=2.0&max-width=1024&background=FFFFFF)
Figure 4: Key HMBC and 1H–1H COSY correlations of 6.
Figure 4: Key HMBC and 1H–1H COSY correlations of 6.
The relative stereochemistry of H-1, H-5, H-6, H-7, Me-14 and Me-15 in 6 was determined by NMR data comparison with literature data and NOESY spectral analysis. The chemical shifts (δC 16.6, 28.7) of the methyl groups of Me-12 and Me-13 correlate well with those previously assigned to the corresponding methyl groups in 8, 9 and other analogues [7-11], which suggests the cis-orientation of the cyclopropane ring [7,8]. The coupling constants (J = 10.0 Hz) between H-5 and H-1/H-6 indicate the trans-relationship of H-5 with H-1 and H-6. By comparison with the known chemical-shift data and shift parameters [7-11], the β-orientation of H-1, H-6, H-7 and α-orientation of H-5 were assigned. In the seven-numbered ring, the methyl group C-15 (δC 17.8) appears at a high field, reflecting that Me-15 is not in the same orientation as H-6 and H-7 [7]. In the NOESY spectrum, correlations of OMe-10 with H-1, H-6, H-7 are observed (Figure 5), which suggest that OMe-10, H-1, H-6, H-7 are in β-orientation, meanwhile, the presence of correlations between H-5 and Me-14/Me-13 indicates that H-5, Me-14, Me-13 are in α-orientation. Thus, the structure of 6 was elucidated to be as shown and named as (+)-4β-N-methenetauryl-10β-methoxy-1β,5α,6β,7β-aromadendrane.
![[1860-5397-8-18-5]](/bjoc/content/figures/1860-5397-8-18-5.png?scale=2.0&max-width=1024&background=FFFFFF)
Figure 5: Key NOESY correlations of 6 and 7.
Figure 5: Key NOESY correlations of 6 and 7.
Compound 7 showed the same molecular formula of C19H35NSO4 as 6 on the basis of NMR spectra and HRMS–ESI (m/z 372.2172 [M − H]−). The 1H and 13C NMR spectral data of 7 (Table 2) showed similarities to those of 6, except for some obvious differences in the chemical shift data of H-5 (from δH 0.98 to δH 1.72), C-3 (from δC 37.0 to δC 33.4), C-5 (from δC 46.0 to δC 41.7), C-9 (from δC 37.2 to δC 32.8), and C-15 (from δC 17.8 to δC 24.4), which were caused by the stereochemistry change. In the 1H–1H COSY spectrum, no correlation between H-1 and H-5 is observed, which suggests a small coupling constant and the cis relationship between H-1 and H-5. The coupling constant (J = 9.5 Hz) between H-5 and H-6 indicates the trans-relationship of H-5 with H-6. In the NOESY spectrum, H-16 shows correlations with H-1/H-5, which suggests that H-16, H-1, and H-5 are in β-orientation; meanwhile, correlations of H-6 with H-7/Me-14/Me-13 indicate that H-6, H-7, Me-14 and Me-13 are in α-orientation. The chemical shift of Me-15 (δC 24.4) appears at a slightly lower field reflecting that Me-15 is at the same α-orientation as H-6 and H-7 [7]. Thus, the structure of 7 was elucidated to be as shown and named as (−)-4β-N-methenetauryl-10β-methoxy-1β,5β,6α,7α-aromadendrane.
The cytotoxicity of compounds 1–9 against human malignant melanoma A735 and cervical carcinoma HeLa cell lines was evaluated by MTT assay. The results show that 1–9 does not exhibit cytotoxicity against A735 and HeLa cell lines with IC50 > 200 μg/mL. Antibacterial activity of compounds 1–5 at a concentration of 25 μg/disc (diameter 6 mm) was measured against bacteria Escherichia coli, Bacillus subtilis, and Micrococcus luteus strains by using standard disc-diffusion assay. The results show that 1–5 exhibit no inhibitory effects towards all tested bacteria at 25 μg/disc, 7 can inhibit the growth of B. subtilis and M. luteus with inhibition zones of 6.3 mm and 6.2 mm, respectively, at 25 μg/disc, while 7 has no effect towards E. coli at 25 μg/disc. It was reported that taurine and its derivatives have a number of physiological functions, including interference with GABA and glycine receptors, antinociceptic effects, anticonvulsive actions, neuroprotective actions, etc. [14]. In this study, because of the limited quantity of compound, we did not further test the other bioactivities of 6 and 7.
Experimental
General experimental procedures
Optical rotations were measured with a Horiba SEAP-300 spectropolarimeter. UV spectra were measured with a Shimadzu double-beam 210A spectrophotometer in MeOH solution. 1H, 13C NMR and 2D NMR spectra were recorded on a Bruker AV-500 MHz NMR spectrometer with TMS as internal standard. MS spectral data were obtained on an LCQDECA XP HPLC/MSn spectrometer for ESIMS. Silica gel (200–300 mesh) for column chromatography and GF254 for TLC were obtained from the Qindao Marine Chemical Factory, Qindao, People’s Republic of China.
Animal material
The South China Sea gorgonian coral Melitodes squamata Nutting was collected in Sanya, Hainan province, China in July 2008 and identified by Prof. Huang H., the South China Sea Institute of Oceanology, Academia Sinica. A voucher specimen (No. 0802) was deposited in the South China Sea Institute of Oceanology, Academia Sinica, Guangzhou, China.
Extraction and isolation
The frozen specimen of the South China Sea gorgonian coral M. squamata (20 kg, wet weight) was extracted with EtOH/CH2Cl2 (2:1) three times at room temperature and the solvent was evaporated in vacuo. The residue was partitioned in H2O and extracted with EtOAc and n-BuOH three times each. The extracts of each respective solvent were combined. The EtOAc and n-BuOH extracts were concentrated in vacuo to afford 35.17 g and 16.85 g of residue, respectively. The EtOAc extract was subjected to silica-gel column chromatography (column A), eluted with CHCl3/MeOH (from 1:10 to 0:10). By combining the fractions with TLC (GF254) monitoring, 17 fractions were obtained. Fraction A-5 was applied to a Sephadex LH-20 column (column B), eluted with CHCl3/MeOH (1:1), to obtain four fractions. Fraction B-4 was further purified by HPLC (MeOH/H2O 45:100) to yield 1 (1.8 mg, tR = 23.5 min), 2 (1.8 mg, tR = 41.2 min), 3 (2.8 mg, tR = 38.2 min), 4 (2.7 mg, tR = 13.8 min) and 5 (1.8 mg, tR = 9.0 min), separately. Fraction B-3 was further purified by RP-18 reverse-phase silica-gel column chromatography (column C), eluted with MeOH/H2O (30:100), to yield 8 (14.8 mg) and 9 (9.5 mg). Fraction A-11 was separated by a Sephadex LH-20 column (column D), eluted with CHCl3/MeOH (1:1) to give five fractions. Fraction D-2 was subjected to silica-gel column chromatography (column E), eluted with CHCl3/MeOH (10:1), to yield 6 (4.2 mg) and three other fractions. Fraction E-2 was furthered purified by a Sephadex LH-20 column (MeOH/H2O 95:5) to yield 7 (4.8 mg).
Obtucarbamate C (1): White powder; UV (MeOH): 258, 216 nm; 1H and 13C NMR data, see Table 1; (+)-ESIMS (m/z): 409.0 [M + Na]+, 795.0 [2M + Na]+; HRMS–ESI (m/z): [M + Na]+ calcd for C19H23N4O5Na, 409.1590; found, 409.1568.
Obtucarbamate D (2): White powder; UV (MeOH): 258, 216 nm; 1H and 13C NMR data, see Table 1; (+)-ESIMS (m/z): 409.1 [M + Na]+, 795.3 [2M + Na]+; HRMS–ESI (m/z): [M + Na]+ calcd for C19H23N4O5Na, 409.1590; found, 409.1576.
(+)-4β-N-methenetauryl-10β-methoxy-1β,5α,6β,7β-aromadendrane (6): White powder; [α]D20 +26.4° (c 0.28, MeOH); 1H and 13C NMR data, see Table 2; (−)-ESIMS (m/z): 372.1 [M − H]−, 745.4 [2M − H]−; HRMS–ESI (m/z): [M − H]− calcd for C19H34NSO4, 372.2209; found, 372.2170.
(−)-4β-N-methenetauryl-10β-methoxy-1β,5β,6α,7α-aromadendrane (7): White powder; [α]D20 −7.5° (c 0.48, MeOH); 1H and 13C NMR data, see Table 2; (−)-ESIMS (m/z): 372.2 [M − H]−, 745.4 [2M − H]−; HRMS–ESI (m/z): [M − H]− calcd for C19H34NSO4, 372.2209; found, 372.2172.
Acknowledgements
The authors are grateful to the National Science Foundation of China (grant 20872151), the research supported by the CAS/SAFEA International Partnership Program for Creative Research Teams (grant KZCX2-YW-T001), the National Basic Research Program of China (grant 2010CB8338-00-G), the National Science Foundation of China (grant 40976090 and 40910093), the Knowledge Innovation Program of Chinese Academy of Science (grant KSCX2-EW-G-12B), and the Hi-tech Research and Development Program of China (grant SQ2007AA09Z409) for financial support.
References
-
Coll, J. C. Chem. Rev. 1992, 92, 613–631. doi:10.1021/cr00012a006
Return to citation in text: [1] -
Rodríguez, A. D. Tetrahedron 1995, 51, 4571–4618. doi:10.1016/0040-4020(95)00216-U
Return to citation in text: [1] -
Shao, C. L.; Fu, X. M.; Wang, C. Y.; Han, L.; Liu, X.; Fang, Y. C.; Li, G. Q.; Zeng, X. Q.; Liu, G. X.; Guan, H. S. Period. Ocean Univ. China 2009, 39, 691–698.
Return to citation in text: [1] -
Kobayashi, M.; Kanda, F. J. Chem. Soc., Perkin Trans. 1 1991, 1177–1179. doi:10.1039/P19910001177
Return to citation in text: [1] -
Brown, G. W.; Cohen, P. P. Biochem. J. 1960, 75, 82–91.
Return to citation in text: [1] [2] [3] [4] [5] [6] -
Kuo, Y. H.; Jou, M. H. Chem. Express 1990, 5, 909–912.
Return to citation in text: [1] [2] [3] [4] [5] [6] -
Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X
Return to citation in text: [1] [2] [3] [4] [5] [6] [7] [8] [9] -
Bohlmann, F.; Grenz, M.; Jakupovic, J.; King, R. M.; Robinson, H. Phytochemistry 1983, 22, 1213–1218. doi:10.1016/0031-9422(83)80224-6
Return to citation in text: [1] [2] [3] [4] [5] -
Beechan, C. M.; Djerassi, C.; Eggert, H. Tetrahedron 1978, 34, 2503–2508. doi:10.1016/0040-4020(78)88378-1
Return to citation in text: [1] [2] [3] [4] -
Jizba, J.; Laudová, V.; Samek, Z.; Ubik, K.; Novotny, L. Collect. Czech. Chem. Commun. 1981, 46, 1048–1053. doi:10.1135/cccc19811048
Return to citation in text: [1] [2] [3] [4] -
Liu, H.-J.; Wua, C.-L.; Becker, H.; Zapp, J. Phytochemistry 2000, 53, 845–849. doi:10.1016/S0031-9422(99)00609-3
Return to citation in text: [1] [2] [3] [4] -
De Marino, S.; Iorizzi, M.; Zollo, F.; Debitus, C.; Menou, J.-L.; Ospina, L. F.; Alcaraz, M. J.; Payá, M. J. Nat. Prod. 2000, 63, 322–326. doi:10.1021/np990374+
Return to citation in text: [1] [2] -
Xynas, R.; Capon, R. J. Aust. J. Chem. 1989, 42, 1427–1433. doi:10.1071/CH9891427
Return to citation in text: [1] [2] -
Simo, S.; Oja, P. S. Proc. West. Pharmacol. Soc. 2007, 50, 8–15.
Return to citation in text: [1]
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 8. | Bohlmann, F.; Grenz, M.; Jakupovic, J.; King, R. M.; Robinson, H. Phytochemistry 1983, 22, 1213–1218. doi:10.1016/0031-9422(83)80224-6 |
| 9. | Beechan, C. M.; Djerassi, C.; Eggert, H. Tetrahedron 1978, 34, 2503–2508. doi:10.1016/0040-4020(78)88378-1 |
| 10. | Jizba, J.; Laudová, V.; Samek, Z.; Ubik, K.; Novotny, L. Collect. Czech. Chem. Commun. 1981, 46, 1048–1053. doi:10.1135/cccc19811048 |
| 11. | Liu, H.-J.; Wua, C.-L.; Becker, H.; Zapp, J. Phytochemistry 2000, 53, 845–849. doi:10.1016/S0031-9422(99)00609-3 |
| 12. | De Marino, S.; Iorizzi, M.; Zollo, F.; Debitus, C.; Menou, J.-L.; Ospina, L. F.; Alcaraz, M. J.; Payá, M. J. Nat. Prod. 2000, 63, 322–326. doi:10.1021/np990374+ |
| 13. | Xynas, R.; Capon, R. J. Aust. J. Chem. 1989, 42, 1427–1433. doi:10.1071/CH9891427 |
| 12. | De Marino, S.; Iorizzi, M.; Zollo, F.; Debitus, C.; Menou, J.-L.; Ospina, L. F.; Alcaraz, M. J.; Payá, M. J. Nat. Prod. 2000, 63, 322–326. doi:10.1021/np990374+ |
| 13. | Xynas, R.; Capon, R. J. Aust. J. Chem. 1989, 42, 1427–1433. doi:10.1071/CH9891427 |
| 1. | Coll, J. C. Chem. Rev. 1992, 92, 613–631. doi:10.1021/cr00012a006 |
| 2. | Rodríguez, A. D. Tetrahedron 1995, 51, 4571–4618. doi:10.1016/0040-4020(95)00216-U |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 8. | Bohlmann, F.; Grenz, M.; Jakupovic, J.; King, R. M.; Robinson, H. Phytochemistry 1983, 22, 1213–1218. doi:10.1016/0031-9422(83)80224-6 |
| 9. | Beechan, C. M.; Djerassi, C.; Eggert, H. Tetrahedron 1978, 34, 2503–2508. doi:10.1016/0040-4020(78)88378-1 |
| 10. | Jizba, J.; Laudová, V.; Samek, Z.; Ubik, K.; Novotny, L. Collect. Czech. Chem. Commun. 1981, 46, 1048–1053. doi:10.1135/cccc19811048 |
| 11. | Liu, H.-J.; Wua, C.-L.; Becker, H.; Zapp, J. Phytochemistry 2000, 53, 845–849. doi:10.1016/S0031-9422(99)00609-3 |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 8. | Bohlmann, F.; Grenz, M.; Jakupovic, J.; King, R. M.; Robinson, H. Phytochemistry 1983, 22, 1213–1218. doi:10.1016/0031-9422(83)80224-6 |
| 9. | Beechan, C. M.; Djerassi, C.; Eggert, H. Tetrahedron 1978, 34, 2503–2508. doi:10.1016/0040-4020(78)88378-1 |
| 10. | Jizba, J.; Laudová, V.; Samek, Z.; Ubik, K.; Novotny, L. Collect. Czech. Chem. Commun. 1981, 46, 1048–1053. doi:10.1135/cccc19811048 |
| 11. | Liu, H.-J.; Wua, C.-L.; Becker, H.; Zapp, J. Phytochemistry 2000, 53, 845–849. doi:10.1016/S0031-9422(99)00609-3 |
| 4. | Kobayashi, M.; Kanda, F. J. Chem. Soc., Perkin Trans. 1 1991, 1177–1179. doi:10.1039/P19910001177 |
| 5. | Brown, G. W.; Cohen, P. P. Biochem. J. 1960, 75, 82–91. |
| 6. | Kuo, Y. H.; Jou, M. H. Chem. Express 1990, 5, 909–912. |
| 3. | Shao, C. L.; Fu, X. M.; Wang, C. Y.; Han, L.; Liu, X.; Fang, Y. C.; Li, G. Q.; Zeng, X. Q.; Liu, G. X.; Guan, H. S. Period. Ocean Univ. China 2009, 39, 691–698. |
| 5. | Brown, G. W.; Cohen, P. P. Biochem. J. 1960, 75, 82–91. |
| 6. | Kuo, Y. H.; Jou, M. H. Chem. Express 1990, 5, 909–912. |
| 5. | Brown, G. W.; Cohen, P. P. Biochem. J. 1960, 75, 82–91. |
| 6. | Kuo, Y. H.; Jou, M. H. Chem. Express 1990, 5, 909–912. |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 8. | Bohlmann, F.; Grenz, M.; Jakupovic, J.; King, R. M.; Robinson, H. Phytochemistry 1983, 22, 1213–1218. doi:10.1016/0031-9422(83)80224-6 |
| 7. | Goldsby, G.; Burke, B. A. Phytochemistry 1987, 26, 1059–1063. doi:10.1016/S0031-9422(00)82350-X |
| 8. | Bohlmann, F.; Grenz, M.; Jakupovic, J.; King, R. M.; Robinson, H. Phytochemistry 1983, 22, 1213–1218. doi:10.1016/0031-9422(83)80224-6 |
| 9. | Beechan, C. M.; Djerassi, C.; Eggert, H. Tetrahedron 1978, 34, 2503–2508. doi:10.1016/0040-4020(78)88378-1 |
| 10. | Jizba, J.; Laudová, V.; Samek, Z.; Ubik, K.; Novotny, L. Collect. Czech. Chem. Commun. 1981, 46, 1048–1053. doi:10.1135/cccc19811048 |
| 11. | Liu, H.-J.; Wua, C.-L.; Becker, H.; Zapp, J. Phytochemistry 2000, 53, 845–849. doi:10.1016/S0031-9422(99)00609-3 |
© 2012 Huang et al; licensee Beilstein-Institut.
This is an Open Access article under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
The license is subject to the Beilstein Journal of Organic Chemistry terms and conditions: (http://www.beilstein-journals.org/bjoc)




![[1860-5397-8-18-2]](/bjoc/content/figures/1860-5397-8-18-2.png?scale=2.0&max-width=1024&background=FFFFFF)