Search results
Search for "glioblastoma" in Full Text gives 16 result(s) in Beilstein Journal of Organic Chemistry.
Anti-invasive and cytotoxic evaluation of a (+)-pinoresinol-based semisynthetic library against glioblastoma
- Chen Zhang,
- Kah Yean Lum,
- Jonathan M. White,
- Paul I. Forster,
- Nicholas Booth,
- Sunita A. Ramesh and
- Rohan A. Davis
Beilstein J. Org. Chem. 2026, 22, 691–704, doi:10.3762/bjoc.22.54
- (+)-pinoresinol and its high abundance from the seeds of E. maculata, we synthesized five analogues for biological evaluations. Preliminary cytotoxicity evaluations of the semisynthetic pinoresinol-based library against two human glioblastoma cell lines, U251MG and KNS42, showed (+)-4,4'-di(3,3-dimethylbutanoyl
- ; glioblastoma; (+)-pinoresinol; seeds; semisynthesis; Introduction Since ancient times, natural products have played a crucial role in the development of medicines for various diseases [1][2][3][4]. A recent review by Newman et al. has identified that over the past four decades, >30% of therapeutic drugs
- cancer cells from moving. In this study, a unique plant-derived lignan‑based library was screened against glioblastoma (brain cancer) cell lines to evaluate anti-invasive and cytotoxicity effects. Results and Discussion The air-dried and ground seeds of E. maculata were sequentially extracted with CH2Cl2

Graphical Abstract

Figure 1: Chemical structures of the two known natural products, (+)-salicifoliol (1) and (+)-pinoresinol (2)...

Figure 2: Reaction schemes detailing the semisynthesis of (+)-eudesmin (3), (+)-phillygenin (4), (+)-5,5'-dib...

Figure 3: Key COSY, HMBC, and ROESY correlations for (+)-5,5'-dibromopinoresinol (5).

Figure 4: ORTEP drawing of (+)-eudesmin (3).

Figure 5: ECD data for compounds 2–7 in MeOH.

Figure 6: Percentage of cell viability for (+)-pinoresinol (2) and its analogues (+)-eudesmin (3), (+)-philly...

Figure 7: (A) Transwell invasive assay of the (+)-pinoresinol library 2–7 on U251MG cells at a concentration ...

Figure 8: (A) Transwell invasion assay of (+)-4,4'-di(3,3-dimethylbutanoyl)pinoresinol (6) on KNS42 cells at ...

Figure 9: The U251 cell line was derived from a grade III–IV malignant tumor, specifically an astrocytoma, ob...
Towards the targeted protein degradation of CK2: design and synthesis of CAM4066-based PROTACs
- Sophie Day-Riley,
- Sona Krajcovicova,
- Aryaman Raj Sokhal,
- Jan L. Venne,
- Paul Brear,
- Marko Hyvönen,
- Benjamin C. Whitehurst,
- Jason S. Carroll and
- David R. Spring
Beilstein J. Org. Chem. 2026, 22, 611–619, doi:10.3762/bjoc.22.47
- survival across several cancer types, and its downregulation impairs viability in models of glioblastoma, medulloblastoma, cholangiocarcinoma, breast and renal cancers [4]. These features have established CK2 as a compelling therapeutic target. In 2016, structural studies revealed a cryptic αD pocket

Graphical Abstract

Figure 1: Design strategy and validation. A) Structure of CAM4066 (1) that served as a model design for the d...

Scheme 1: Synthesis of the key ligand-linker PROTAC precursors. For compound 14, HATU/DIPEA has been used ins...

Scheme 2: (A) Synthesis of final VHL PROTACs; (B) Synthesis of final CRBN PROTACs. Abbreviations: Boc = tert-...

Figure 2: Full structures of final VHL PROTACs 23–26 and CRBN PROTACs 28 and 29.
Design, synthesis and biological evaluation of 2,5-diaryloxazolo[4,5-d]pyrimidin-7-ylamines as selective cytotoxic agents against HeLa cells
- Maryna V. Kachaeva,
- Agnieszka B. Olejniczak,
- Marta Denel-Bobrowska,
- Victor V. Zhirnov,
- Yevheniia S. Velihina,
- Stepan G. Pilyo and
- Volodymyr S. Brovarets
Beilstein J. Org. Chem. 2026, 22, 390–398, doi:10.3762/bjoc.22.27
- lines such as HepG2 (liver), HeLa (cervix), A549 (lung), and glioblastoma models rely on enhanced nucleotide metabolism to sustain rapid DNA/RNA synthesis. Oxazolopyrimidines can inhibit these overactive enzymes in cancer cells, while normal cells (with lower demand) are less affected. This explains why
- carcinoma cells), and T98G (Human glioblastoma multiforme cells). Cytotoxicity was determined by measurement of 50% inhibition of cell growth by the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay. The selectivity index (SI) was calculated for the investigated compounds. The
- (ATCC CCL-185, Human lung carcinoma cells), HepG2 (ATCC HB-8065, Human hepatocellular carcinoma cells), and T98G (ATCC CRL-1690, Human glioblastoma multiforme cells). Non-cancer cell line: NCTC clone 929 (CCL-1, Mus musculus normal subcutaneous connective tissue cells). Chemicals: Eagles minimum

Graphical Abstract

Scheme 1: Rational design of [1,3]oxazolo[4,5-d]pyrimidin-7-amine derivatives 1–9.

Scheme 2: Synthesis of new 2,5-diaryloxazolo[4,5-d]pyrimidin-7-ylamines 1–9.
N-Glycosides of indigo, indirubin, and isoindigo: blue, red, and yellow sugars and their cancerostatic activity
- Peter Langer
Beilstein J. Org. Chem. 2024, 20, 2840–2869, doi:10.3762/bjoc.20.240
- cytotoxic activity against melanoma and squamous cell carcinoma cells, lung cancer and glioblastoma cells [30]. It was shown in a WST-1 assay that these compounds exhibit a concentration-dependent inhibition of the metabolic activity of A375 and A431 cells, while the non-glycosylated derivative 34 proved to

Graphical Abstract

Scheme 1: Structures of indigo (1a), indirubin (2a) and isoindigo (3a).

Scheme 2: Structures of akashins A–C.

Scheme 3: Synthesis of 5b. Reagents and conditions: i) TMSOTf, 4 Å MS, CH2Cl2, −20 °C, 1.5 h, then 20 °C, 8–1...

Scheme 4: Synthesis of 7c. Reagents and conditions: i) TMSOTf, 4 Å MS, CH2Cl2, −18 °C, 3 h; then: TMSOTf, 4 Å...

Scheme 5: Synthesis of 1d. Reagents and conditions: i) chloroacetic acid, Na2CO3, reflux, 6 h; ii) Ac2O, NaOA...

Scheme 6: Synthesis of 10e. Reagents and conditions: i) p-TsOH·H2O, acetonitrile, MeOH, 1 d; ii) NIS, PPh3, D...

Scheme 7: Synthesis of akashins A–C. Reagents and conditions: i) TMSOTf, 4 Å MS, CH2Cl2, −18 to 20 °C, 15 h; ...

Scheme 8: Synthesis of 5d. Reagents and conditions: i) KMnO4, AcOH, high-power-stirring (12.000 rot/min), 20 ...

Scheme 9: Possible mechanism of the formation of 5c.

Scheme 10: Synthesis of 7d. Reagents and conditions: i) 1) CH2Cl2, 2) Me3SiI, 20 °C, 30 min, 3) 0 °C, 30 min, ...

Scheme 11: Synthesis of α-15b. Reagents and conditions: i) 1) CH2Cl2, 2) Me3SiI, 20 °C, 30 min, 3) 0 °C, 30 mi...

Scheme 12: Synthesis of isatin-N-glycosides 16a–f. Reagents and conditions: i) PhNH2, EtOH, 20 °C, 12 h; ii) Ac...

Scheme 13: Synthesis of 17–21. Reagents and conditions: i) Na2CO3, MeOH, 20 °C, 4 h.

Scheme 14: Synthesis of indirubin-N-glycosides α-17a and α-17b.

Scheme 15: Synthesis of β-17f. Reagents and conditions: i) 1) Na2CO3, MeOH, 20 °C, 4 h, 2) Ac2O/pyridine 1:1, ...

Scheme 16: Synthesis of β-24a. Reagents and conditions: i) n-PrOH, H2O, formic acid (buffer, 100 mM), 2 h, 65 ...

Scheme 17: Synthesis of isatin-N-glycosides 23b–g and 24b–g.

Scheme 18: Synthesis of β-29a,b. Reagents and conditions: i) EtOH, 20 °C, 12 h; ii) DDQ, dioxane, 20 °C, 12 h;...

Scheme 19: Synthesis of β-31a. Reagents and conditions: i) Na2SO3, dioxane, H2O, 110 °C, 2 d; ii) piperidine, ...

Scheme 20: Synthesis of 33a–d. Reagents and conditions: i) Ac2O, AcOH, NaOAc, 80 °C, 1 h; ii) 1) NaOMe, anhydr...

Scheme 21: Indirubins 34 and 35.

Scheme 22: Synthesis of 36f. Reagents and conditions: i) NaOH, H2O, 20 °C, 5 h; ii) HCl, NaNO2, H2O, −14 °C; i...

Scheme 23: Synthesis of 38a–h. Reagents and conditions: i) 1) 0.1 equiv NaOMe, MeOH, 20 °C, 15–20 min, 2) HOAc...

Scheme 24: Synthesis of 40a–h. Reagents and conditions: i) method A: EtOH/THF, cat. KOt-Bu, 20 °C, 3–4.5 h; me...

Scheme 25: Synthesis of 41a–d. Reagents and conditions: i) Ac2O, AcOH, NaOAc, 80 °C, 1 h.

Scheme 26: Synthesis of 41e. Reagents and conditions: i) AcOH, NaOAc, 110 °C, 24 h.

Scheme 27: Synthesis of E-β-43a–e and E-β-44a,b. Reagents and conditions: i) 1) NEt3, EtOH, 20 °C, 12 h, 2) DM...

Scheme 28: Synthesis of E-43f. Reagents and conditions: i) Na2CO3, MeOH, 20 °C, 6–24 h.

Scheme 29: Synthesis of 46a–m. Reagents and conditions: i) NEt3 (1 equiv), EtOH, 20 °C, 6–10 h; ii) MsCl, NEt3...

Scheme 30: Synthesis of 48a–d. Reagents and conditions: i) AcOH/Ac2O, NaOAc, 60 °C, 3–4 h.

Scheme 31: Synthesis of 48e. Reagents and conditions: i) NaOAc, AcOH, 110 °C, 24 h.

Scheme 32: Synthesis of β-49a,b. Reagents and conditions: i) AcOH/Ac2O, NaOAc, 60 °C, 3–4 h.

Scheme 33: Synthesis of β-54a,b. Reagents and conditions: i) 1) NaH, DMF, 0 °C, 15 min, 2) β-51a,b, 20 °C, 3 h...

Scheme 34: Synthesis of 54c–l. The yields refer to the yields of the first and second condensation step for ea...

Scheme 35: Synthesis of 57a–c and 58a–d. Reagents and conditions: i) HCl (conc.), AcOH, reflux, 24 h; ii) 1) B...

Scheme 36: Synthesis of 59a–e and 60a–e. Reagents and conditions: i) P(NEt2)3 (1.1 equiv), CH2Cl2, −78 °C to 2...

Scheme 37: Synthesis of 61a–d and 62a–d. Reagents and conditions: i) P(NEt2)3 (1.1 equiv), CH2Cl2, −78 °C to 2...

Scheme 38: Synthesis of β-64a–e and α-64a. Reagents and conditions: i) AcOH, Ac2O, NaOAc, 90 °C, 6 h.

Scheme 39: Synthesis of β-72a. Reagents and conditions: i) 66, EtOH, 20 °C, 12 h; ii) DDQ, dioxane, 20 °C, 12 ...

Scheme 40: Synthesis of β-72b.

Scheme 41: Synthesis of β-74a–c. Reagents and conditions: i) AcOH, Ac2O, NaOAc, 130 °C, 2 d.

Scheme 42: Synthesis of β-77. Reagents and conditions: i) 1) NEt3, EtOH, 20 °C, 12 h, 2) DMAP, NEt3, MsCl, 0 °...

Scheme 43: Synthesis of β-81a–f and β-80g. Reagents and conditions: i) AcOH, 80 °C, 1–3 h; ii) benzene, PTSA, ...

Scheme 44: Synthesis of 84a. Reagents and conditions: i) benzene, AlCl3, 20 °C, 10 min; ii) MeOH, NaOMe, 12 h,...

Scheme 45: Synthesis of 84b–l. The yields refer to the yields of the condensation and the deprotection step fo...
The Groebke–Blackburn–Bienaymé reaction in its maturity: innovation and improvements since its 21st birthday (2019–2023)
- Cristina Martini,
- Muhammad Idham Darussalam Mardjan and
- Andrea Basso
Beilstein J. Org. Chem. 2024, 20, 1839–1879, doi:10.3762/bjoc.20.162


Graphical Abstract

Scheme 1: Mechanism of the GBB reaction.

Scheme 2: Comparison of the performance of Sc(OTf)3 with some RE(OTf)3 in a model GBB reaction. Conditions: a...

Scheme 3: Comparison of the performance of various Brønsted acid catalysts in the synthesis of GBB adduct 6. ...

Scheme 4: Synthesis of Brønsted acidic ionic liquid catalyst 7. Conditions: a) neat, 60 °C, 24 h; b) TfOH, DC...

Scheme 5: Aryliodonium derivatives as organic catalysts in the GBB reaction. In the box the proposed binding ...

Scheme 6: DNA-encoded GBB reaction in micelles made of amphiphilic polymer 13. Conditions: a) 13 (50 equiv), ...

Scheme 7: GBB reaction catalyzed by cyclodextrin derivative 14. Conditions: a) 14 (1 mol %), water, 100 °C, 4...

Scheme 8: Proposed mode of activation of CALB. a) activation of the substrates; b) activation of the imine; c...

Scheme 9: One-pot GBB reaction–Suzuki coupling with a bifunctional hybrid biocatalyst. Conditions: a) Pd(0)-C...

Scheme 10: GBB reaction employing 5-HMF (23) as carbonyl component. Conditions: a) TFA (20 mol %), EtOH, 60 °C...

Scheme 11: GBB reaction with β-C-glucopyranosyl aldehyde 26. Conditions: a) InCl3 (20 mol %), MeOH, 70 °C, 2–3...

Scheme 12: GBB reaction with diacetylated 5-formyldeoxyuridine 29, followed by deacetylation of GBB adduct 30....

Scheme 13: GBB reaction with glycal aldehydes 32. Conditions: a) HFIP, 25 °C, 2–4 h.

Scheme 14: Vilsmeier–Haack formylation of 6-β-acetoxyvouacapane (34) and subsequent GBB reaction. Conditions: ...

Scheme 15: GBB reaction of 4-formlyl-PCP 37. Conditions: a) HOAc or HClO4, MeOH/DCM (2:3), rt, 3 d.

Scheme 16: GBB reaction with HexT-aldehyde 39. Conditions: a) 39 (20 nmol) and amidine (20 μmol), MeOH, rt, 6 ...

Scheme 17: GBB reaction of 2,4-diaminopirimidine 41. Conditions: a) Sc(OTf)3 (20 mol %), MeCN, 120 °C (MW), 1 ...

Scheme 18: Synthesis of N-edited guanine derivatives from 3,6-diamine-1,2,4-triazin-5-one 44. Conditions: a) S...

Scheme 19: Synthesis of 2-aminoimidazoles 49 by a Mannich-3CR followed by a one-pot intramolecular oxidative a...

Scheme 20: On DNA Suzuki–Miyaura reaction followed by GBB reaction. Conditions: a) CsOH, sSPhos-Pd-G2; b) AcOH...

Scheme 21: One-pot cascade synthesis of 5-iminoimidazoles. Conditions: a) Na2SO4, DMF, 220 °C (MW).

Scheme 22: GBB reaction of 5-amino-1H-imidazole-4-carbonile 57. Conditions: a) HClO4 (5 mol %), MeOH, rt, 24 h....

Scheme 23: One-pot cascade synthesis of indole-imidazo[1,2,a]pyridine hybrids. In blue the structural motif in...

Scheme 24: One-pot cascade synthesis of fused polycyclic indoles 67 or 69 from indole-3-carbaldehyde. Conditio...

Scheme 25: One-pot cascade synthesis of linked- and bridged polycyclic indoles from indole-2-carbaldehyde (70)...

Scheme 26: One-pot cascade synthesis of pentacyclic dihydroisoquinolines (X = N or CH). In blue the structural...

Scheme 27: One-pot stepwise synthesis of imidazopyridine-fused benzodiazepines 85. Conditions: a) p-TsOH (20 m...

Scheme 28: One-pot stepwise synthesis of benzoxazepinium-fused imidazothiazoles 89. Conditions: a) Yb(OTf)3 (2...

Scheme 29: One-pot stepwise synthesis of fused imidazo[4,5,b]pyridines 95. Conditions: a) HClO4, MeOH, rt, ove...

Scheme 30: Synthesis of heterocyclic polymers via the GBB reaction. Conditions: a) p-TsOH, EtOH, 70 °C, 24 h.

Scheme 31: One-pot multicomponent reaction towards the synthesis of covalent organic frameworks via the GBB re...

Scheme 32: One-pot multicomponent reaction towards the synthesis of covalent organic frameworks via the GBB re...

Scheme 33: GBB-like multicomponent reaction towards the synthesis of benzothiazolpyrroles (X = S) and benzoxaz...

Scheme 34: GBB-like multicomponent reaction towards the formation of imidazo[1,2,a]pyridines. Conditions: a) I2...

Scheme 35: Post-functionalization of GBB products via Ugi reaction. Conditions a) HClO4, DMF, rt, 24 h; b) MeO...

Scheme 36: Post-functionalization of GBB products via Click reaction. Conditions: a) solvent-free, 150 °C, 24 ...

Scheme 37: Post-functionalization of GBB products via cascade alkyne–allene isomerization–intramolecular nucle...

Scheme 38: Post-functionalization of GBB products via metal-catalyzed intramolecular N-arylation. In red and b...

Scheme 39: Post-functionalization of GBB products via isocyanide insertion (X = N or CH). Conditions: a) HClO4...

Scheme 40: Post-functionalization of GBB products via intramolecular nucleophilic addition to nitriles. Condit...

Scheme 41: Post-functionalization of GBB products via Pictet–Spengler cyclization. Conditions: a) 4 N HCl/diox...

Scheme 42: Post-functionalization of GBB products via O-alkylation. Conditions: a) TFA (20 mol %), EtOH, 120 °...

Scheme 43: Post-functionalization of GBB products via macrocyclization (X = -CH2CH2O-, -CH2-, -(CH2)4-). Condi...

Figure 1: Antibacterial activity of GBB-Ugi adducts 113 on both Gram-negative and Gram-positive strains.

Scheme 44: GBB multicomponent reaction using trimethoprim as the precursor. Conditions: a) Yb(OTf)3 or Y(OTf)3...

Figure 2: Antibacterial activity of GBB adducts 152 against MRSA and VRE; NA = not available.

Figure 3: Antibacterial activity of GBB adduct 153 against Leishmania amazonensis promastigotes and amastigot...

Figure 4: Antiviral and anticancer evaluation of the GBB adducts 154a and 154b. In vitro antiproliferative ac...

Figure 5: Anticancer activity of the GBB-furoxan hybrids 145b, 145c and 145d determined through antiprolifera...

Scheme 45: Synthesis and anticancer activity of the GBB-gossypol conjugates. Conditions: a) Sc(OTf)3 (10 mol %...

Figure 6: Anticancer activity of polyheterocycles 133a and 136a against human neuroblastoma. Clonogenic assay...

Figure 7: Development of GBB-adducts 158a and 158b as PD-L1 antagonists. HTRF assays were carried out against...

Figure 8: Development of imidazo[1,2-a]pyridines and imidazo[1,2-a]pyrazines as TDP1 inhibitors. The SMM meth...

Figure 9: GBB adducts 164a–c as anticancer through in vitro HDACs inhibition assays. Additional cytotoxic ass...

Figure 10: GBB adducts 165, 166a and 166b as anti-inflammatory agents through HDAC6 inhibition; NA = not avail...

Scheme 46: GBB reaction of triphenylamine 167. Conditions: a) NH4Cl (10 mol %), MeOH, 80 °C (MW), 1 h.

Scheme 47: 1) Modified GBB-3CR. Conditions: a) TMSCN (1.0 equiv), Sc(OTf)3 (0.2 equiv), MeOH, 140 °C (MW), 20 ...

Scheme 48: GBB reaction to assemble imidazo-fused heterocycle dimers 172. Conditions: a) Sc(OTf)3 (20 mol %), ...

Figure 11: Model compounds 173 and 174, used to study the acid/base-triggered reversible fluorescence response...
Cycloaddition reactions of heterocyclic azides with 2-cyanoacetamidines as a new route to C,N-diheteroarylcarbamidines
- Pavel S. Silaichev,
- Tetyana V. Beryozkina,
- Vsevolod V. Melekhin,
- Valeriy O. Filimonov,
- Andrey N. Maslivets,
- Vladimir G. Ilkin,
- Wim Dehaen and
- Vasiliy A. Bakulev
Beilstein J. Org. Chem. 2024, 20, 17–24, doi:10.3762/bjoc.20.3
- embryo kidney cells (HEK-293), glioblastoma (A-172), and osteosarcoma (HOS) cells. Supporting Information Supporting Information File 25: Full experimental details and characterization data of all new compounds. Supporting Information File 26: Copies of NMR spectra of all new compounds. Funding Design

Graphical Abstract

Scheme 1: Synthesis of heteroaryl amidines.

Figure 1: Structures of starting compounds.

Scheme 2: Scope of 3,3-diaminoacrylonitriles 1 and heterocyclic azides 2. Reaction conditions: 1 (0.5 mmol), 2...

Scheme 3: Proposed mechanism for the formation of triazoles 3.
Efficient N-arylation of 4-chloroquinazolines en route to novel 4-anilinoquinazolines as potential anticancer agents
- Rodolfo H. V. Nishimura,
- Thiago dos Santos,
- Valter E. Murie,
- Luciana C. Furtado,
- Leticia V. Costa-Lotufo and
- Giuliano C. Clososki
Beilstein J. Org. Chem. 2021, 17, 2968–2975, doi:10.3762/bjoc.17.206
- adenocarcinoma), and T98G (human glioblastoma). Initially, we evaluated all compounds at 50 µM, and we considered active the compounds that inhibited cell proliferation by over 75% for each cell line. Compounds 10b, 10c, 10g, 10k, 10l, 15a, 15b, and 15d were active against T98G cells, and compounds 10b, 15a, and
- strongly cytotoxic to T98G cells (0.2 nM). In fact, the promising activity of verubulin in glioma is known, which has prompted phase 2 clinical trials in patients with recurrent glioblastoma, but the studies have been interrupted due to the observed adverse events [33]. Nonetheless, further studies aiming

Graphical Abstract

Figure 1: Some antitumor agents containing the 4-anilinoquinazoline moiety.

Scheme 1: Examples of N-arylation reactions using 4-chloroquinazolines as substrates.

Scheme 2: Synthesis of verubulin analog.

Scheme 3: Synthesis of 4-chloro-6-halo-2-phenylquinazolines 8a and 8b. Conditions: a) NBS, CH3CN, 30 min, 25 ...

Scheme 4: N-Arylation reactions using ortho-, meta-, and para-substituted primary anilines of type 14 followe...

Scheme 5: N-Arylation reactions using 4-chloroquinazoline (16) and 4-chloro-2-methylquinazoline (17) to achie...
Synthesis and anticancer activity of bis(2-arylimidazo[1,2-a]pyridin-3-yl) selenides and diselenides: the copper-catalyzed tandem C–H selenation of 2-arylimidazo[1,2-a]pyridine with selenium
- Mio Matsumura,
- Tsutomu Takahashi,
- Hikari Yamauchi,
- Shunsuke Sakuma,
- Yukako Hayashi,
- Tadashi Hyodo,
- Tohru Obata,
- Kentaro Yamaguchi,
- Yasuyuki Fujiwara and
- Shuji Yasuike
Beilstein J. Org. Chem. 2020, 16, 1075–1083, doi:10.3762/bjoc.16.94

- Figure 6, 2f also exhibited cytotoxicity against U251 human glioblastoma cells and HKBMM human malignant meningioma cells. Importantly, the diselenide 2f was more cytotoxic toward HeLa cancer cells than to human brain microvascular endothelial (HBME) cells, which are noncancerous (Figure 7). Hence, the
- represented as mean ± S.D. values of four samples. *p < 0.05; **p < 0.01 compared to the control. DOX: doxorubicin. The cytotoxic effect of the bis[2-(4-methoxyphenyl)imidazo[1,2-a]pyridin-3-yl] diselenide 2f on cancer cell lines. Human glioblastoma U251 cells (a) and human malignant meningioma HKBMM cells (b

Graphical Abstract

Figure 1: Biologically active selenides and diselenides having heteroaryl groups.

Figure 2: Ortep drawings of 2a (a and b) and 3a (c and d, thermal elipsoids indicate 50% probability).

Figure 3: The synthesis of bis(2-arylimidazopyridin-3-yl) diselenides. Reaction conditions: 1 (2 mmol), Se (2...

Figure 4: The synthesis of bis(2-arylimidazopyridin-3-yl) selenides. Reaction conditions: 1 (2 mmol), Se (1 m...

Scheme 1: Control reactions.

Scheme 2: Proposed mechanism (1).

Scheme 3: Proposed mechanism (2).

Figure 5: The cytotoxic effect of the bis(2-arylimidazo[1,2-a]pyridin-3-yl) diselenides 2 and selenides 3 on ...

Figure 6: The cytotoxic effect of the bis[2-(4-methoxyphenyl)imidazo[1,2-a]pyridin-3-yl] diselenide 2f on can...

Figure 7: Cytotoxic effect of the bis[2-(4-methoxyphenyl)imidazo[1,2-a]pyridin-3-yl] diselenide 2f on a cance...
Fluorinated phenylalanines: synthesis and pharmaceutical applications
- Laila F. Awad and
- Mohammed Salah Ayoup
Beilstein J. Org. Chem. 2020, 16, 1022–1050, doi:10.3762/bjoc.16.91

- further analysis and comparison with 178 (performed in vitro), as the most promising candidate 2-[18F]-2-fluoroethyl-ʟ-phenylalanine (2-[18F]FELP, 95) was selected. In a F98 glioblastoma rat model, 2-[18F]FELP exhibited improved in vitro characteristics over [18F]FET 178, especially in view of the

Graphical Abstract

Figure 1: Categories I–V of fluorinated phenylalanines.

Scheme 1: Synthesis of fluorinated phenylalanines via Jackson’s method.

Scheme 2: Synthesis of all-cis-tetrafluorocyclohexylphenylalanines.

Scheme 3: Synthesis of ʟ-4-[sulfono(difluoromethyl)]phenylalanine (nPt: neopentyl, TCE: trichloroethyl).

Scheme 4: Synthesis of ʟ-4-[sulfono(difluoromethyl)]phenylalanine derivatives 17.

Scheme 5: Synthesis of fluorinated Phe analogues from Cbz-protected aminomalonates.

Scheme 6: Synthesis of tetrafluorophenylalanine analogues via the 3-methyl-4-imidazolidinone auxiliary 25.

Scheme 7: Synthesis of tetrafluoro-Phe derivatives via chiral auxiliary 31.

Scheme 8: Synthesis of 2,5-difluoro-Phe and 2,4,5-trifluoro-Phe via Schöllkopf reagent 34.

Scheme 9: Synthesis of 2-fluoro- and 2,6-difluoro Fmoc-Phe derivatives starting from chiral auxiliary 39.

Scheme 10: Synthesis of 2-[18F]FPhe via chiral auxiliary 43.

Scheme 11: Synthesis of FPhe 49a via photooxidative cyanation.

Scheme 12: Synthesis of FPhe derivatives via Erlenmeyer azalactone synthesis.

Scheme 13: Synthesis of (R)- and (S)-2,5-difluoro Phe via the azalactone method.

Scheme 14: Synthesis of 3-bromo-4-fluoro-(S)-Phe (65).

Scheme 15: Synthesis of [18F]FPhe via radiofluorination of phenylalanine with [18F]F2 or [18F]AcOF.

Scheme 16: Synthesis of 4-borono-2-[18F]FPhe.

Scheme 17: Synthesis of protected 4-[18F]FPhe via arylstannane derivatives.

Scheme 18: Synthesis of FPhe derivatives via intermediate imine formation.

Scheme 19: Synthesis of FPhe derivatives via Knoevenagel condensation.

Scheme 20: Synthesis of FPhe derivatives 88a,b from aspartic acid derivatives.

Scheme 21: Synthesis of 2-(2-fluoroethyl)phenylalanine derivatives 93 and 95.

Scheme 22: Synthesis of FPhe derivatives via Zn2+ complexes.

Scheme 23: Synthesis of FPhe derivatives via Ni2+ complexes.

Scheme 24: Synthesis of 3,4,5-trifluorophenylalanine hydrochloride (109).

Scheme 25: Synthesis of FPhe derivatives via phenylalanine aminomutase (PAM).

Scheme 26: Synthesis of (R)-2,5-difluorophenylalanine 115.

Scheme 27: Synthesis of β-fluorophenylalanine via 2-amino-1,3-diol derivatives.

Scheme 28: Synthesis of β-fluorophenylalanine derivatives via the oxazolidinone chiral auxiliary 122.

Scheme 29: Synthesis of β-fluorophenylalanine from pyruvate hemiketal 130.

Scheme 30: Synthesis of β-fluorophenylalanine (136) via fluorination of β-hydroxyphenylalanine (137).

Scheme 31: Synthesis of β-fluorophenylalanine from aziridine derivatives.

Scheme 32: Synthesis of β-fluorophenylalanine 136 via direct fluorination of pyruvate esters.

Scheme 33: Synthesis of β-fluorophenylalanine via fluorination of ethyl 3-phenylpyruvate enol using DAST.

Scheme 34: Synthesis of β-fluorophenylalanine derivatives using photosensitizer TCB.

Scheme 35: Synthesis of β-fluorophenylalanine derivatives using Selectflour and dibenzosuberenone.

Scheme 36: Synthesis of protected β-fluorophenylalanine via aziridinium intermediate 150.

Scheme 37: Synthesis of β-fluorophenylalanine derivatives via fluorination of α-hydroxy-β-aminophenylalanine d...

Scheme 38: Synthesis of β-fluorophenylalanine derivatives from α- or β-hydroxy esters 152a and 155.

Scheme 39: Synthesis of a series of β-fluoro-Phe derivatives via Pd-catalyzed direct fluorination of β-methyle...

Scheme 40: Synthesis of series of β-fluorinated Phe derivatives using quinoline-based ligand 162 in the Pd-cat...

Scheme 41: Synthesis of β,β-difluorophenylalanine derivatives from 2,2-difluoroacetaldehyde derivatives 164a,b....

Scheme 42: Synthesis of β,β-difluorophenylalanine derivatives via an imine chiral auxiliary.

Scheme 43: Synthesis of α-fluorophenylalanine derivatives via direct fluorination of protected Phe 174.

Figure 2: Structures of PET radiotracers of 18FPhe derivatives.

Figure 3: Structures of melfufen (179) and melphalan (180) anticancer drugs.

Figure 4: Structure of gastrazole (JB95008, 181), a CCK2 receptor antagonist.

Figure 5: Dual CCK1/CCK2 antagonist 182.

Figure 6: Structure of sitagliptin (183), an antidiabetic drug.

Figure 7: Structure of retaglpitin (184) and antidiabetic drug.

Figure 8: Structure of evogliptin (185), an antidiabetic drug.

Figure 9: Structure of LY2497282 (186) a DPP-4 inhibitor for the treatment of type II diabetes.

Figure 10: Structure of ulimorelin (187).

Figure 11: Structure of GLP1R (188).

Figure 12: Structures of Nav1.7 blockers 189 and 190.
Lanyamycin, a macrolide antibiotic from Sorangium cellulosum, strain Soce 481 (Myxobacteria)
- Lucky S. Mulwa,
- Rolf Jansen,
- Dimas F. Praditya,
- Kathrin I. Mohr,
- Patrick W. Okanya,
- Joachim Wink,
- Eike Steinmann and
- Marc Stadler
Beilstein J. Org. Chem. 2018, 14, 1554–1562, doi:10.3762/bjoc.14.132
- (KB3.1), colon carcinoma cells (HCT-116), and glioblastoma (U87MG). It exhibited cytotoxic activity with IC50 values of 3.1 and 1.5 µM against L929 and KB3.1 cell cultures, respectively, while it was not active against the other cell lines according to the limits of the National Cancer Institute (active

Graphical Abstract

Figure 1: Lanyamycin (1/2), differing at C-26, isolated from Sorangium cellulosum (Soce 481).

Figure 2: Structure fragments of lanyamycin (1/2) from 1H,1H-COSY spectrum (bold bonds) and selected 1H,13C-H...

Figure 3: Structure of bafilomycin A1.

Figure 4: 3D Model of the macrolactone ring of lanyamycin (1/2) and selected ROESY correlations. Green arrows...

Figure 5: HCV assay: NC = negative control, EGCG = positive control, Cpd1 = lanyamycin (1/2).
An overview of recent advances in duplex DNA recognition by small molecules
- Sayantan Bhaduri,
- Nihar Ranjan and
- Dev P. Arya
Beilstein J. Org. Chem. 2018, 14, 1051–1086, doi:10.3762/bjoc.14.93
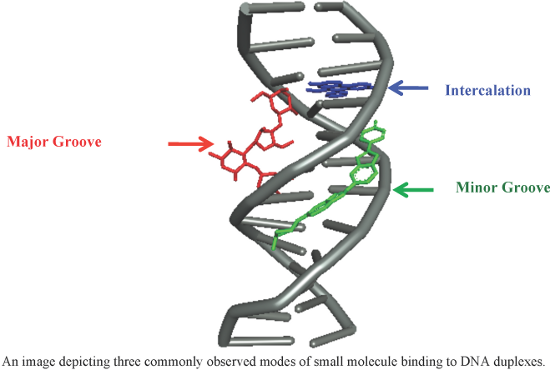

Graphical Abstract

Figure 1: A figure showing the hydrogen bonding patterns observed in (a) duplex (b) triplex and (c) quadruple...

Figure 2: (a) Portions of MATα1–MATα2 are shown contacting the minor groove of the DNA substrate. Key arginin...

Figure 3: Chemical structures of naturally occurring and synthetic hybrid minor groove binders.

Figure 4: Synthetic structural analogs of distamycin A by replacing one or more pyrrole rings with other hete...

Figure 5: Pictorial representation of the binding model of pyrrole–imidazole (Py/Im) polyamides based on the ...

Figure 6: Chemical structures of synthetic “hairpin” pyrrole–imidazole (Py/Im) conjugates.

Figure 7: (a) Minor groove complex formation between DNA duplex and 8-ring cyclic Py/Im polyamide (conjugate ...

Figure 8: Telomere-targeting tandem hairpin Py/Im polyamides 23 and 24 capable of recognizing >10 base pairs; ...

Figure 9: Representative examples of recently developed DNA minor groove binders.

Figure 10: Chemical structures of bisbenzamidazoles Hoechst 33258 and 33342 and their synthetic structural ana...

Figure 11: Chemical structures of bisamidines such as diminazene, DAPI, pentamidine and their synthetic struct...

Figure 12: Representative examples of recently developed bisamidine derivatives.

Figure 13: Chemical structures of chromomycin, mithramycin and their synthetic structural analogs 91 and 92.

Figure 14: Chemical structures of well-known naturally occurring DNA binding intercalators.

Figure 15: Naturally occurring indolocarbazole rebeccamycin and its synthetic analogs.

Figure 16: Representative examples of naturally occurring and synthetic derivatives of DNA intercalating agent...

Figure 17: Several recent synthetic varieties of DNA intercalators.

Figure 18: Aminoglycoside (neomycin)–Hoechst 33258/intercalator conjugates.

Figure 19: Chemical structures of triazole linked neomycin dimers and neomycin–bisbenzimidazole conjugates.

Figure 20: Representative examples of naturally occurring and synthetic analogs of DNA binding alkylating agen...

Figure 21: Chemical structures of naturally occurring and synthetic analogs of pyrrolobenzodiazepines.
On the design principles of peptide–drug conjugates for targeted drug delivery to the malignant tumor site
- Eirinaios I. Vrettos,
- Gábor Mező and
- Andreas G. Tzakos
Beilstein J. Org. Chem. 2018, 14, 930–954, doi:10.3762/bjoc.14.80

- receptor-mediated transcytosis after binding to LRP-1 and consequently it is often used as drug delivery vehicle, while paclitaxel bears cytotoxicity against glioblastoma. It has been shown that the brain uptake of ANG1005 was 4.5-fold higher compared to paclitaxel and the cytotoxicity remained higher in
- all cancer cell lines tested (glioblastoma U87 MG, U118, U251; lung carcinoma A549, NCI-H460, Calu-3; ovarian carcinoma SK-OV-3). It has been also proved that human tumor xenografts were inhibited more with ANG1005 than paclitaxel. Finally, mice with intracerebral implantation of U87 MG glioblastoma

Graphical Abstract

Figure 1: Conventional chemotherapy versus targeted chemotherapy. Black color = Solid malignant tumor; red = ...

Figure 2: A. General structural architecture of the ideal navigated drug delivery system [31]. B. General structu...

Figure 3: Binding and penetration mechanism of iRGD. The iRGD peptide is accumulated on the surface of αv int...

Figure 4: Representative examples of anticancer drugs utilized for the construction of PDCs. The most usual c...

Figure 5: Illustration of the drug release mechanism from the self-immolative spacer PABC conjugated to a tum...

Figure 6: Structures of the PDCs named AN-152 and AN-207.

Figure 7: Structure of the PDC named AN-238.

Figure 8: Chemical structure and synthetic scheme for the PDC ANG1005. (A) ANG1005 is composed of three molec...

Figure 9: Structure of oxime linked Dau–GnRH-III conjugate with or without cathepsin B labile spacer and thei...

Figure 10: Synthesis of the most effective GnRH-III–Dau conjugate with two drug molecules [153].

Figure 11: Structures of the four different PDCs of D-Lys6-GnRH-I and gemcitabine (GSG, GSG2, 3G, 3G2) [19].

Figure 12: Structures of (A) native sunitinib; (B) SAN1 analog of sunitinib and (C) assembled PDC named SAN1GS...

Figure 13: Synthetic scheme for the formation of GSG and the unexpected side product [156].

Figure 14: Illustration of uncharted guanidinium peptide coupling reagent side reactions during PDCs synthesis ...

Figure 15: Putative mechanism for the formation of the uronium side product [156].
Synthesis and biological evaluation of RGD and isoDGR peptidomimetic-α-amanitin conjugates for tumor-targeting
- Lizeth Bodero,
- Paula López Rivas,
- Barbara Korsak,
- Torsten Hechler,
- Andreas Pahl,
- Christoph Müller,
- Daniela Arosio,
- Luca Pignataro,
- Cesare Gennari and
- Umberto Piarulli
Beilstein J. Org. Chem. 2018, 14, 407–415, doi:10.3762/bjoc.14.29
- antiproliferative activity of the conjugates was evaluated in three cell lines possessing different levels of αVβ3 integrin expression: human glioblastoma U87 (αVβ3+), human lung carcinoma A549 (αVβ3−) and breast adenocarcinoma MDA-MB-468 (αVβ3−). In the U87, in the MDA-MB-468, and partly in the A549 cancer cell
- disulfide linker in the cytoplasm. In another example, Perrin and co-workers conjugated the N-propargylasparagine of an amanitin analog to a cycloRGD integrin ligand (cyclo[RGDfK]) using a copper-catalyzed azide–alkyne cycloaddition [8]. The conjugates were tested in the U87 glioblastoma cell line, but only
- a slight enhancement in toxicity over α-amanitin was observed. The transmembrane receptor αVβ3 integrin is widely expressed on the blood vessels of several human cancers (for example, breast cancer, glioblastoma, pancreatic tumor, prostate carcinoma) but not on the vasculature of healthy tissues [9

Graphical Abstract

Figure 1: α-Amanitin.

Figure 2: Structure of the ligands cyclo[DKP-RGD] (1), NH2CH2-cyclo[DKP-RGD] (2), cyclo[DKP-isoDGR] (3), NH2CH...

Figure 3: Structure of the α-amanitin conjugates: cyclo[DKP-RGD]-uncleavable-α-amanitin (7), cyclo[DKP-isoDGR...

Scheme 1: Synthesis of α-amanitin-cyclo[DKP-RGD] and α-amanitin-cyclo[DKP-isoDGR] conjugates 7–10. Reagents a...

Scheme 2: Synthesis of cyclo[DKP-isoDGR]-PEG-4-Val-Ala-α-amanitin conjugate 11. Reagents and conditions: a) 4...
C-5’-Triazolyl-2’-oxa-3’-aza-4’a-carbanucleosides: Synthesis and biological evaluation
- Roberto Romeo,
- Caterina Carnovale,
- Salvatore V. Giofrè,
- Maria A. Chiacchio,
- Adriana Garozzo,
- Emanuele Amata,
- Giovanni Romeo and
- Ugo Chiacchio
Beilstein J. Org. Chem. 2015, 11, 328–334, doi:10.3762/bjoc.11.38
- ]: Some of these compounds show a good anticancer activity against the anaplastic (8305C) and the follicular (FTC-133) human thyroid cancer cell lines, and especially on the U87MG human primary glioblastoma cell line (Figure 2). Accordingly, considering that the incorporation of the triazole moiety can

Graphical Abstract

Figure 1: 2’-Oxa-3’-aza-modified nucleosides and 2’-oxa-3’-aza-modified nucleotides.

Figure 2: Triazolyl-2’-oxa-3’-aza-4’a-carbanucleosides.

Scheme 1: Synthesis of triazolyl isoxazolidinyl-nucleosides 13 and 14. Reagents and conditions: a) Tosyl chlo...

Figure 3: α–β Epimerization.
An easily accessible sulfated saccharide mimetic inhibits in vitro human tumor cell adhesion and angiogenesis of vascular endothelial cells
- Grazia Marano,
- Claas Gronewold,
- Martin Frank,
- Anette Merling,
- Christian Kliem,
- Sandra Sauer,
- Manfred Wiessler,
- Eva Frei and
- Reinhard Schwartz-Albiez
Beilstein J. Org. Chem. 2012, 8, 787–803, doi:10.3762/bjoc.8.89
- angiogenesis and is currently under investigation in several clinical trials for the treatment of recurrent malignant glioblastoma [48]. GSF, due to its direct anti-tumor and anti-angiogenic effect, may represent a new class of small-molecule anticancer drugs. Although anti-angiogenic drugs such as the

Graphical Abstract

Scheme 1: Synthesis of (4-{[(β-D-galactopyranosyl)oxy]methyl}furan-3-yl)methyl hydrogen sulfate (GSF, 5) and ...

Figure 1: Effects of increasing concentrations of (4-{[(β-D-galactopyranosyl)oxy]methyl}furan-3-yl)methyl hyd...

Figure 2: Inhibition of adhesion of WM-115 cells to fibrinogen (A), or to fibronectin (B) with increasing con...

Figure 3: Inhibition of adhesion of melanoma cells WM-115 to fibronectin-coated plastic by 5 mM (4-{[(β-D-gal...

Figure 4: In silico blind-docking (A, B) and molecular dynamic simulations (C) of (4-{[(β-D-galactopyranosyl)...

Figure 5: Intact cell monolayers of WM-115 cells in 12-well plates were wounded with a 100 µL pipette tip and...

Figure 6: A: Zymograms (color inverted) of serum-free conditioned medium of melanoma cells treated with (4-{[...

Figure 7: Adhesion of HBMEC-60 to extracellular matrix proteins. Prior to the adhesion experiments, HBMEC-60 ...

Figure 8: Effect of (4-{[(β-D-galactopyranosyl)oxy]methyl}furan-3-yl)methyl hydrogen sulfate (GSF) on transmi...

Figure 9: Influence of saccharide mimetics on endothelial networking (matrigel-assay) (A) and tube formation ...
Recent progress on the total synthesis of acetogenins from Annonaceae
- Nianguang Li,
- Zhihao Shi,
- Yuping Tang,
- Jianwei Chen and
- Xiang Li
Beilstein J. Org. Chem. 2008, 4, No. 48, doi:10.3762/bjoc.4.48

Graphical Abstract

Scheme 1: Total synthesis of longifolicin by Marshall’s group.

Scheme 2: Total synthesis of corossoline by Tanaka’s group.

Scheme 3: Total synthesis of corossoline by Wu’s group.

Scheme 4: Total synthesis of pseudo-annonacin A by Hanessian’s group.

Scheme 5: Total synthesis of tonkinecin by Wu’s group.

Scheme 6: Total synthesis of gigantetrocin A by Shi’s group.

Scheme 7: Total synthesis of annonacin by Wu’s group.

Scheme 8: Total synthesis of solamin by Kitahara’s group.

Scheme 9: Total synthesis of solamin by Mioskowski’s group.

Scheme 10: Total synthesis of cis-solamin by Makabe’s group.

Scheme 11: Total synthesis of cis-solamin by Brown’s group.

Scheme 12: The formal synthesis of (+)-cis-solamin by Donohoe’s group.

Scheme 13: Total synthesis of cis-solamin by Stark’s group.

Scheme 14: Total synthesis of mosin B by Tanaka’s group.

Scheme 15: Total synthesis of longicin by Hanessian’s group.

Scheme 16: Total synthesis of murisolin and 16,19-cis-murisolin by Tanaka’s group.

Scheme 17: Synthesis of a stereoisomer library of (+)-murisolin by Curran’s group.

Scheme 18: Total synthesis of murisolin by Makabe’s group.

Scheme 19: Total synthesis of reticulatain-1 by Makabe’s group.

Scheme 20: Total synthesis of muricatetrocin C by Ley’s group.

Scheme 21: Total synthesis of (4R,12S,15S,16S,19R,20R,34S)-muricatetrocin (146) and (4R,12R,15S,16S,19R,20R,34S...

Scheme 22: Total synthesis of parviflorin by Hoye’s group.

Scheme 23: Total synthesis of parviflorin by Trost’s group.

Scheme 24: Total synthesis of trilobacin by Sinha’s group.

Scheme 25: Total synthesis of 15-epi-annonin I 181b by Scharf’s group.

Scheme 26: Total synthesis of squamocin A and squamocin D by Scharf’s group.

Scheme 27: Total synthesis of asiminocin by Marshall’s group.

Scheme 28: Total synthesis of asiminecin by Marshall’s group.

Scheme 29: Total synthesis of (+)-(30S)-bullanin by Marshall’s group.

Scheme 30: Total synthesis of uvaricin by the group of Sinha and Keinan.

Scheme 31: Formal synthesis of uvaricin by Burke’s group.

Scheme 32: Total synthesis of trilobin by Marshall’s group.

Scheme 33: Total synthesis of trilobin by the group of Sinha and Keinan.

Scheme 34: Total synthesis of asimilobin by the group of Wang and Shi.

Scheme 35: Total synthesis of squamotacin by the group of Sinha and Keinan.

Scheme 36: Total synthesis of asimicin by Marshall’s group.

Scheme 37: Total synthesis of asimicin by the group of Sinha and Keinan.

Scheme 38: Total synthesis of asimicin by Roush’s group.

Scheme 39: Total synthesis of asimicin by Marshall’s group.

Scheme 40: Total synthesis of 10-hydroxyasimicin by Ley’s group.

Scheme 41: Total synthesis of asimin by Marshall’s group.

Scheme 42: Total synthesis of bullatacin by the group of Sinha and Keinan.

Scheme 43: Total synthesis of bullatacin by Roush’s group.

Scheme 44: Total synthesis of bullatacin by Pagenkopf’s group.

Scheme 45: Total synthesis of rollidecins C and D by the group of Sinha and Keinan.

Scheme 46: Total synthesis of 30(S)-hydroxybullatacin by Marshall’s group.

Scheme 47: Total synthesis of uvarigrandin A and 5(R)-uvarigrandin A by Marshall’s group.

Scheme 48: Total synthesis of membranacin by Brown’s group.

Scheme 49: Total synthesis of membranacin by Lee’s group.

Scheme 50: Total synthesis of rolliniastatin 1 and rollimembrin by Lee’s group.

Scheme 51: Total synthesis of longimicin D by the group of Maezaki and Tanaka.

Scheme 52: Total synthesis of the structure proposed for mucoxin by Borhan’s group.

Scheme 53: Modular synthesis of adjacent bis-THF annonaceous acetogenins by Marshall’s group.

Scheme 54: Total synthesis of 4-deoxygigantecin by Tanaka’s group.

Scheme 55: Total synthesis of squamostatins D by Marshall’s group.

Scheme 56: Total synthesis of gigantecin by Crimmins’s group.

Scheme 57: Total synthesis of gigantecin by Hoye’s group.

Scheme 58: Total synthesis of cis-sylvaticin by Donohoe’s group.

Scheme 59: Total synthesis of 17(S),18(S)-goniocin by Sinha’s group.

Scheme 60: Total synthesis of goniocin and cyclogoniodenin T by the group of Sinha and Keinan.

Scheme 61: Total synthesis of jimenezin by Takahashi’s group.

Scheme 62: Total synthesis of jimenezin by Lee’s group.

Scheme 63: Total synthesis of jimenezin by Hoffmann’s group.

Scheme 64: Total synthesis of muconin by Jacobsen’s group.

Scheme 65: Total synthesis of (+)-muconin by Kitahara’s group.

Scheme 66: Total synthesis of muconin by Takahashi’s group.

Scheme 67: Total synthesis of muconin by the group of Yoshimitsu and Nagaoka.

Scheme 68: Total synthesis of mucocin by the group of Sinha and Keinan.

Scheme 69: Total synthesis of mucocin by Takahashi’s group.

Scheme 70: Total synthesis of (−)-mucocin by Koert’s group.

Scheme 71: Total synthesis of mucocin by the group of Takahashi and Nakata.

Scheme 72: Total synthesis of mucocin by Evans’s group.

Scheme 73: Total synthesis of mucocin by Mootoo’s group.

Scheme 74: Total synthesis of (−)-mucocin by Crimmins’s group.

Scheme 75: Total synthesis of pyranicin by the group of Takahashi and Nakata.

Scheme 76: Total synthesis of pyranicin by Rein’s group.

Scheme 77: Total synthesis of proposed pyragonicin by the group of Takahashi and Nakata.

Scheme 78: Total synthesis of pyragonicin by Rein’s group.

Scheme 79: Total synthesis of pyragonicin by Takahashi’s group.

Scheme 80: Total synthesis of squamostanal A by Figadère’s group.

Scheme 81: Total synthesis of diepomuricanin by Tanaka’s group.

Scheme 82: Total synthesis of (−)-muricatacin [(R,R)-373a] and its enantiomer (+)-muricatacin [(S,S)-373b] by ...

Scheme 83: Total synthesis of epi-muricatacin (+)-(S,R)-373c and (−)-(R,S)-373d by Scharf’s group.

Scheme 84: Total synthesis of (−)-muricatacin 373a and 5-epi-(−)-muricatacin 373d by Uang’s group.

Scheme 85: Total synthesis of four stereoisomers of muricatacin by Yoon’s group.

Scheme 86: Total synthesis of (+)-muricatacin by Figadère’s group.

Scheme 87: Total synthesis of (+)-epi-muricatacin and (−)-muricatacin by Couladouros’s group.

Scheme 88: Total synthesis of muricatacin by Trost’s group.

Scheme 89: Total synthesis of (−)-(4R,5R)-muricatacin by Heck and Mioskowski’s group.

Scheme 90: Total synthesis of muricatacin (−)-373a by the group of Carda and Marco.

Scheme 91: Total synthesis of (−)- and (+)-muricatacin by Popsavin’s group.

Scheme 92: Total synthesis of (−)-muricatacin by the group of Bernard and Piras.

Scheme 93: Total synthesis of (−)-muricatacin by the group of Yoshimitsu and Nagaoka.

Scheme 94: Total synthesis of (−)-muricatacin by Quinn’s group.

Scheme 95: Total synthesis of montecristin by Brückner’s group.

Scheme 96: Total synthesis of (−)-acaterin by the group of Franck and Figadère.

Scheme 97: Total synthesis of (−)-acaterin by Singh’s group.

Scheme 98: Total synthesis of (−)-acaterin by Kumar’s group.

Scheme 99: Total synthesis of rollicosin by Quinn’s group.

Scheme 100: Total synthesis of Rollicosin by Makabe’s group.

Scheme 101: Total synthesis of squamostolide by Makabe’s group.

Scheme 102: Total synthesis of tonkinelin by Makabe’s group.






